9. Ontology mapping with Ontology Xref Service (OxO)¶

9.1. Main Objectives¶
Different ontology terms can describe the same concept, which makes it difficult for data integration.
Ontology mapping, or ontology alignment, is the process of determining correspondences between equivalent
concepts in alternative ontologies and distinct vocabularies.
OxO maps terms in different ontologies, vocabularies and coding standards using evidence collected mainly from the Ontology Lookup Service (OLS), Unified Medical Language System (UMLS), but also from other sources.
This recipe shows how to use the EMBL-EBI Ontology Xref Service (OxO) service to map ontology terms between source and target vocabularies 1 .
9.3. Requirements¶
recipe dependency:
knowledge requirements:
Familiar with metadata curation
Familiar with ontology and other vocabularies
technical requirements:
Experience with API or Bash scripts allows users to try the automated solutions.
9.4. FAIRification Objectives, Inputs and Outputs¶
Actions.Objectives.Tasks |
Input |
Output |
|---|---|---|
9.5. Table of Data Standards¶
Data Formats |
Terminologies |
Models |
|---|---|---|
9.6. Using EMBL-EBI OXO¶
Ontology cross Ontology or OxO service is available from a web interface or a programmatic interface. The following sections show both.
9.6.1. OXO Web Interface:¶
9.6.1.1. Step 1: Identify source vocabulary and target vocabulary¶
For pathology data annotation, vocabularies, such as MeSH, NCI thesaurus, ICD-10, UMLS, Human Disease Ontology(DOID), and MONDO disease ontology, are widely used. In this example, we use MeSH and DOID as source vocabularies and MONDO as the target vocabulary to demonstrate the ontology mapping workflow.
Obviously, a target vocabulary needs to be selected before mapping. A variety of factors can influence the selection of a terminology resource for a specific purpose or curation context. For more information about this topic, more detailed guidance on how to select vocabularies can be found here.
OxO allows users to explore the mapping status between the target vocabulary and other vocabularies. Figure 1 shows how terms in MONDO are linked to terms in other ontologies. MONDO has over 16,000 mappings to terms in UMLS. It is also mapped to terms in DOID, Ophanet, OMIM, MeSH, etc.
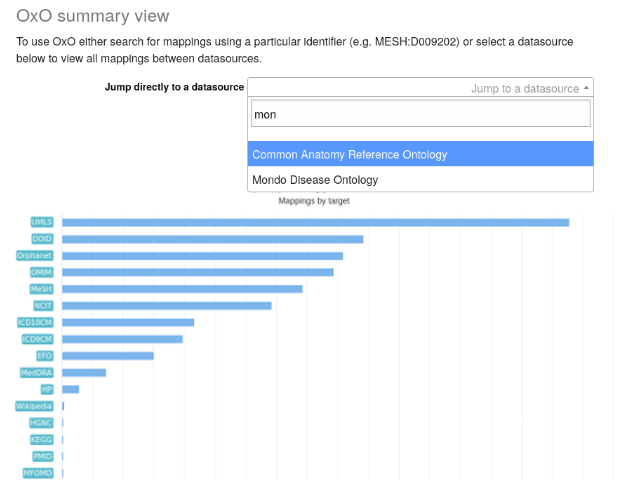
Fig. 9.13 OxO summary result overview.¶
9.6.2. Step 2: Provide the source vocabulary term ID¶
OxO takes ontology term IDs as inputs for ontology mapping, which assumes the (meta)data has been annotated. If the data is not annotated, please check the ontology annotation recipe first.
In this example, we use terms for type 2 diabetes from 2 distinct sources. The corresponding term IDs are listed in the table below.
Text |
Corresponding term in MeSH |
Corresponding term in DOID |
|---|---|---|
type 2 diabetes |
9.6.3. Step 3: Perform mapping¶
Figure 2 shows how to map the MeSH and DOID terms to the MONDO disease ontology. Users are expected to:
specify the target ontology
provide a list of source term IDs
indicate the expected mapping distance.
In this example, we set the mapping distance to 1, which uses high confidence mapping evidence only.
Having greater mapping distance returns more mappings but decreases the mapping confidence.
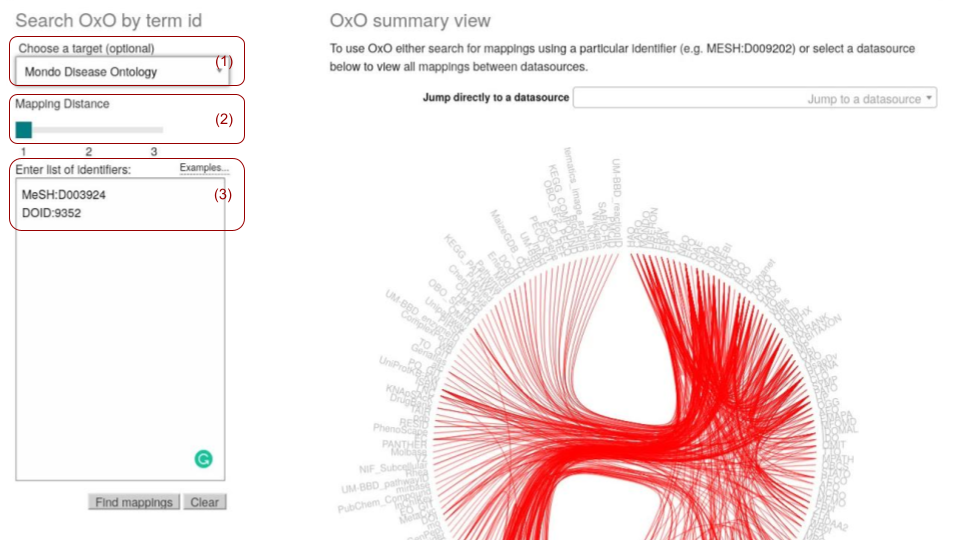
Fig. 9.14 OxO mapping with direct user inputs¶
Figure 3 shows the corresponding mapping results. Both terms have been mapped to ‘MONDO:0005148’. The results can be downloaded as flat-table(tsv) files.
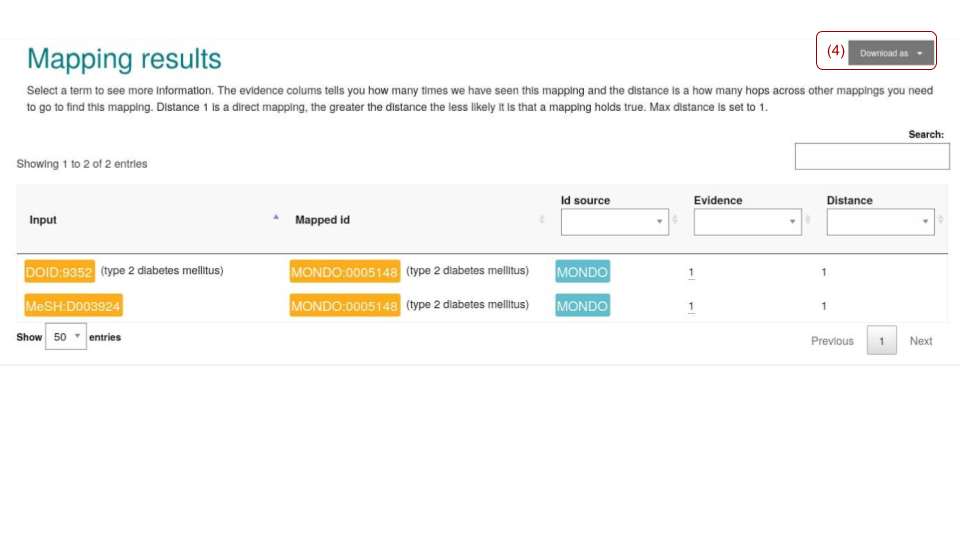
Fig. 9.15 OxO mapping results tabular view.¶
9.6.4. Step 4: Review mapping results¶
To ensure the quality of the mapping, users are recommended to review the mapping results, especially when the mapping distance increases. OxO allows users to explore vocabulary term relationships (see figure 4) by providing a network view of all linked terms. Users can also find more information about each term in the Ontology lookup service.
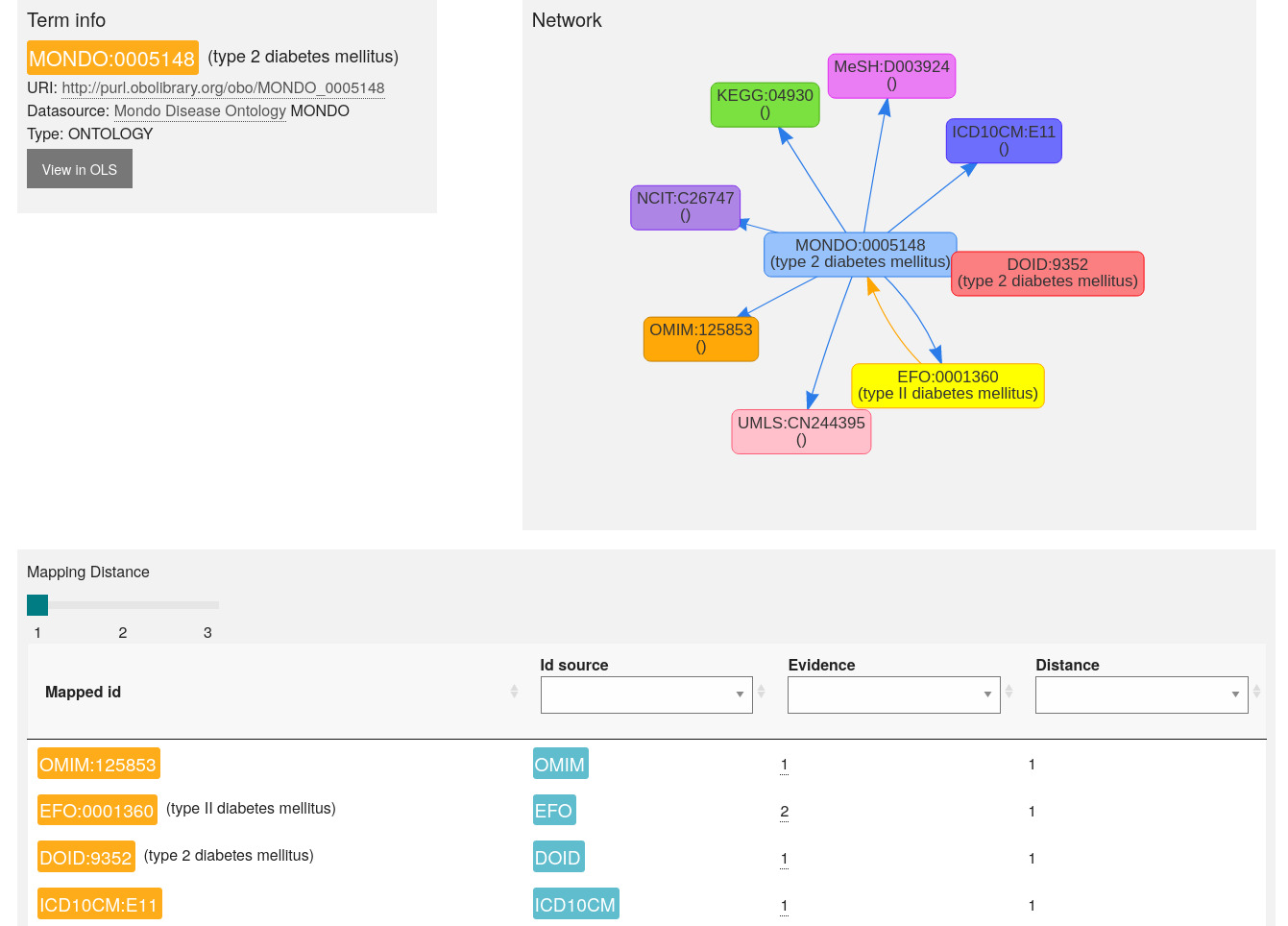
Fig. 9.16 OxO mapping results review - MONDO example.¶
9.7. Using OxO service programmatically¶
Besides the graphical user interface offered by OxO at EMBL-EBI, a REST style API is also available. It is therefore possible to invoke the mapping service from the command line. See the OxO API documentation page.
In the following code snippet, an API request invoked via curl performed the same mapping presented in the section above, at step 3:
curl 'https://www.ebi.ac.uk/spot/oxo/api/search' -i
-X POST
-H 'Content-Type: application/json'
-H 'Accept: application/json'
-d '{
"ids" : [ "DOID:9352","MeSH:D003924"],
"inputSource" : null,
"mappingTarget" : [ "MONDO" ],
"mappingSource" : [ "MONDO" ],
"distance" : 1
}'
The corresponding results are as follows:
{
"queryId" : "DOID:9352",
"querySource" : null,
"curie" : "DOID:9352",
"label" : "type 2 diabetes mellitus",
"mappingResponseList" : [ {
"curie" : "MONDO:0005148",
"label" : "type 2 diabetes mellitus",
"sourcePrefixes" : [ "MONDO" ],
"targetPrefix" : "MONDO",
"distance" : 1
} ],
"_links" : {
"self" : {
"href" : "https://www.ebi.ac.uk/spot/oxo/api/terms/DOID:9352"
},
"mappings" : {
"href" : "https://www.ebi.ac.uk/spot/oxo/api/mappings?fromId=DOID:9352"
}
}
}
9.7.1. Common questions about OxO¶
What about incorrect mappings?
OxO imports term mapping information from ontologies and other curated databases. It doesn’t validate the mapping evidence. Users are recommended to check the mapping results manually, especially when the mapping distance is above 1.
Why is no mapping found?
OxO relies on the ontology and other curated databases to improve the mapping coverage. Some terms describing the same concept might not be aligned in OxO. In this case, users are recommended to identify possible mappings by searching in OLS.
To help the ontology community improve the mapping, users can also submit an update request in corresponding ontologies. The guidance can be found here Requesting new terms
Ontology mapping service for internal databases
OxO has been dockerized for local deployment and licensed under Apache License 2.0. The source code can be found here.
9.8. Conclusion¶
This recipe presented OxO, EMBL-EBI tool for performing ontology mapping as an example to demonstrate the ontology mapping workflow. In the context of a FAIRification workflow, a tool such as OxO comes particurlary handy to augment a dataset with cross-references or replace an annotation set with another in a data integration exercice for instance.
9.8.1. What to read next?¶

FAIRsharing records appearing in this recipe:
9.9. References¶
References
- 1
Oxo is a service for finding mappings (or cross-references) between terms from ontologies, vocabularies and coding standards. 2021. URL: https://www.ebi.ac.uk/spot/oxo/.
9.10. Authors¶
Authors
Name |
ORCID |
Affiliation |
Type |
ELIXIR Node |
Contribution |
|---|---|---|---|---|---|
EMBL-EBI |
|
Writing - Original Draft |
|||
Karsten Quast |
Boehringer-Ingelheim AG |
Writing - Original Draft |
|||
GSK |
Writing - Original Draft |
||||
University of Oxford |
Writing - Review & Editing |


